RO-Crate Bioimaging profiles (detached)#
RO-Crate Bioimaging profiles are being developed as a machine-readable exchange mechanism for microscopy data.
A first iteration of experimentation with RO-Crates for bioimaging happened in the context of the OME 2024 NGFF Challenge. For that, a minimal design was proposed and used for managing contextual metadata on heterogeneous datasets.
The details of the use of RO-Crates in the context of the challenge, as well as a few other uses directly related to the Open Microscopy Environment are reported as an use case on the RO-Crate Website.
Similarly, a detached RO-Crate profile, heavily inspired by the MICrate here, is being developed as an interoperability mechanism for study-level metadata as part of the FoundingGIDE project. The GIDE project does not connect RO-Crate to entry points for the data, but to the pages for studies on data repositories, such as BioImage Archive and the Image Data Resource.
MICrate Specification#
follow up on ro-crate bioimaging minimal design
Version v1.0.0-draft
detached RO-Crate
Authors: Susanne Kunis
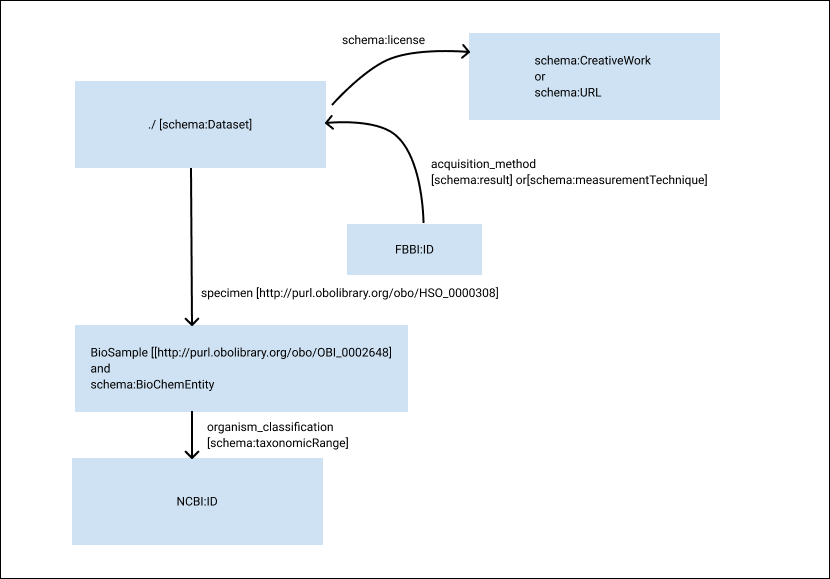
RO-Crate Metadata Descriptor#
{
"@id": "ro-crate-metadata.json",
"@type": "CreativeWork",
"conformsTo": {"@id": "https://w3id.org/ro/crate/1.2"},
"about": {"@id": <url_zarr>}
}
Root Data Entity#
-
@id:URL(MUST)@type:Dataset(MUST)name:Text(SHOULD)
[identify the dataset to humans well enough to disambiguate it from other RO-Crates]description:Text(SHOULD)
[further elaborate on the name to provide a summary of the context in which the dataset is important]license:CreativeWorkorURLorText(SHOULD)
[link to a Contextual Entity in the RO-Crate Metadata File with a name and description. MAY have a URI (eg for Creative Commons or Open Source licenses). MAY, if necessary be a textual description of how the RO-Crate may be used]acquisition_method:DefinedTermorURL(COULD)
[link to Contextual Entity in the Ro-Crate Metadata File describing the method with name and linked specimen. MAY have a termCode and a inDefinedTermSet (eg if Contextual Entity is of type DefinedTerm).]specimen:BioChemEntity(MUST)
[to specify this Ro-Crate is about this particular subject]hasPart:CreativeWorkor subtype of this (COULD)
Contextual Entity#
-
@id:TextorURL(MUST)
[MUST be either a URI Path relative to the RO Crate root, or an absolute URI.]@type:DefinedTermorURL(MUST),inDefinedTermSet: (http://purl.obolibrary.org/obo/) (COULD)
-
@id:TextorURL(MUST)
[MUST be either a URI Path relative to the RO Crate root, or an absolute URI.]@type:BioChemEntity(MUST)organism_classification:DefinedTermorTaxonorTextorURL(MUST)
[organism classification via id or via link to Contextual Entity in the Ro-Crate Metadata File describing the organism. MAY have a name to present the human readable name of the term.]
Custom Context and Extended Metadata:#
Allows to define custom fields within your RO-Crate metadata document and describe how they should be interpreted.
"@context": [
"https://w3id.org/ro/crate/1.2/context",
{
"organism_classification": "https://schema.org/taxonomicRange",
"obo": "http://purl.obolibrary.org/obo/",
"acquisiton_method": {
"@reverse": "https://schema.org/result",
"@type": "@id"
},
"biosample": "http://purl.obolibrary.org/obo/OBI_0002648",
"specimen": "http://purl.obolibrary.org/obo/HSO_0000308"
}
],
Minimal example#
See mic-ro-crate-metadata.json open in:
